Modeling myosin mechanobiology across scales
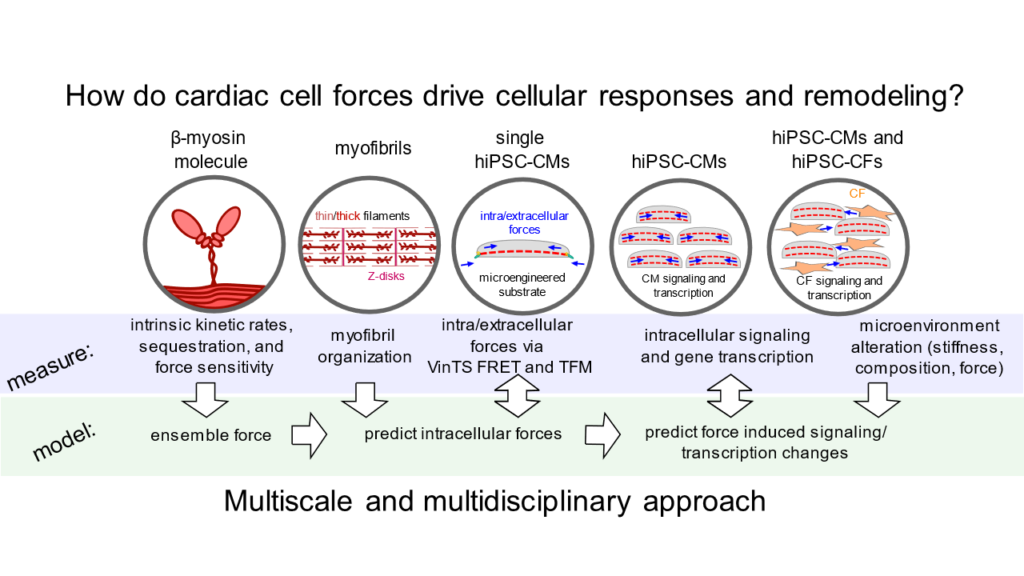
This project focuses on understanding how mutations in beta cardiac myosin with diverse and often opposing effects on myosin molecular function contribute to similar disease phenotypes of hypertrophic cardiomyopathy (HCM): cardiomyocyte (CM) hypertrophy, hypercontractility, and tissue fibrosis. Understanding disease mechanisms and the heterogeneity of phenotypes across multiple scales (molecular, cellular, and clinical) will enable the development of better and more individualized therapies for patients with HCM.
- The first aim of this proposal is to clarify the mechanisms by which altered intracellular forces affect intracellular signaling and CM hypertrophy.
- The second aim is to use computational modeling to examine how alterations to interrelated parameters of myosin function change force production temporally and spatially.
- The third aim is to define how altered extracellular mechanics affect CM and cardiac fibroblast phenotypes.
The project uses several innovative molecular, cellular, bioengineering, and computational modeling tools to examine and contextualize the role of altered cellular mechanics in driving changes in cardiac cell phenotypes. CRISPR/Cas9 gene editing in human induced pluripotent stem cells provides a model system to investigate the effects of specific mutations in a dynamic cellular context. Micropatterned engineered platforms will be used to improve myofibril alignment and allow direct measurement of intracellular force production by traction force microscopy.
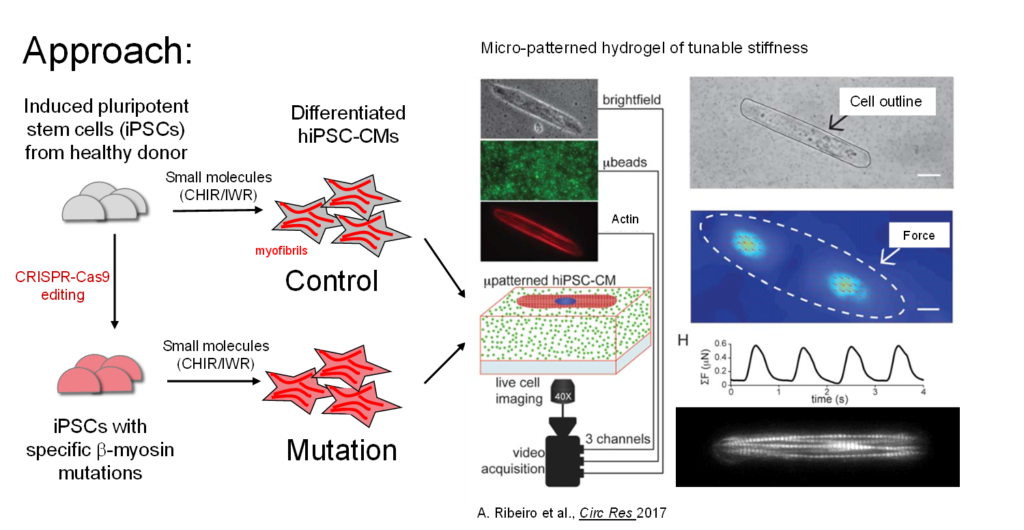
Molecular biology and transcriptomic techniques will be used to measure changes in intracellular signaling and gene expression in response to mechanical perturbation. Another innovation is the novel application of a vinculin tension sensor FRET probe to directly measure intracellular forces at sarcomere Z disks and at cellular adhesions.
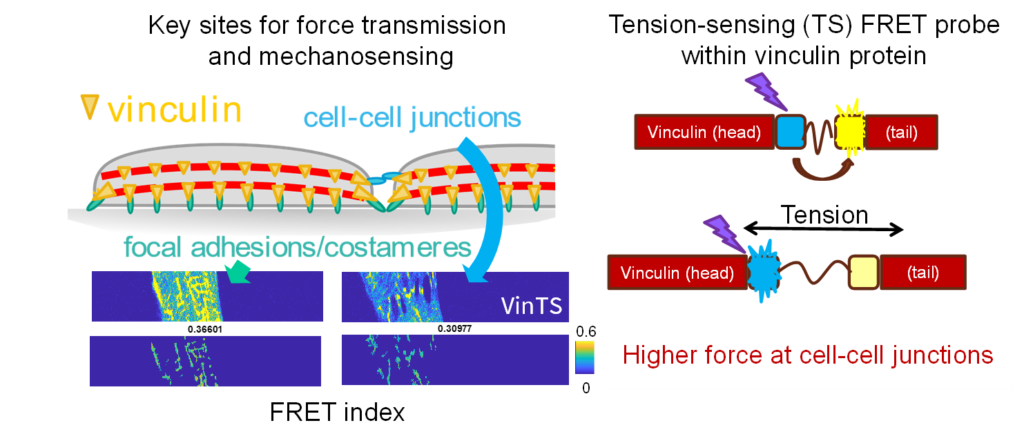
Together these platforms allow direct validation of temporal and spatial cellular forces predicted using computational modeling (molecular myosin models and cell-specific finite element models). Finally, investigating the effect of altered extracellular mechanics on CM function and cardiac fibroblast activation in engineered environments will give insights into the effects of fibrotic remodeling.
Additional developing projects
- Quantitatively linking sarcomeric contraction and force generation to energy (ATP) consumption and cellular metabolism
- Assessing hiPSC-CM population heterogeneity and the impact of mixtures of myosin isoforms
- Extending cellular scale mechanics to organ/cardiovascular system level mechanics and hemodynamics via computational modeling
- Understanding impacts of sex differences/reproductive hormones and pregnancy on cardiac remodeling and hypertrophy (with M-BRISC)
